Rapid evolution of selfing syndrome in plants studied with resurrection ecology
Introduction
The resurrection method is a new ecological approach for the study of rapid adaptation in natural populations (Franks et al. 2018). A basic resurrection study is a common garden experiment where ancestral genotypes (preserved as seeds in natural seed banks or seed repositories) are grown side-by-side with their descendant genotypes (sampled in the same site at present). This setting allows for instant comparison between ancestral descendant genotypes. If phenotypic trait shifts are detected between ancestors and descendants, they can be interpreted as the outcome of rapid evolution.
The resurrection method opens the possibility for the study of rapid evolution in arable weeds, which are exposed to new, intense selective pressures with the development of modern agriculture and global anthropogenic changes. In particular, pollinator decline, coupled with phenological mismatches in flowering time due to temperature fluctuations, could lead towards increased self-pollination and a decreased attractivity of plants to pollinators (selfing syndrome). Alternatively, arable weeds could evolve towards increasing their attractivity to pollinators, thus maintaining pollinator interactions despite the reduction in pollination services. Empirical observation of contemporary evolution in arable weeds has produced data in line with both hypotheses (Brys and Jacquemyn 2012; Thomann et al. 2015; Valencia-Montoya et al. 2021; Cheptou et al. 2022; Acoca-Pidolle et al. 2024), and it is likely that the direction of trait changes is species- and even population-specific.
I use the resurrection method to investigate the effects of climate change and pollinator decline on the evolution of reproductive and pollination strategies in entomophilous arable weeds. The research was funded by several small grants (British Ecological Society, Johanna Quandt Young Academy) and connects three research institutions (University of Ostrava, Goethe-University of Frankfurt, University of Rennes) through joint supervision of several MSc and BSc students that have been working on three independent studies. All of the studies have shown phenotypic trait shifts over very short evolutionary times (17-28 years). Moreover, each study has implemented an additional innovative approach by combining the resurrection method with other eco-evolutionary and molecular tools.
Resurrection ecology with reciprocal transplants “in time”
To reliably assess whether phenotypic trait shifts between ancestors and descendants are adaptive, their fitness needs to be assessed in a reciprocal transplant experiment. Given that actual transplants in time are an impossible feat to achieve, I simulated past- and present-day conditions in climate chamber by regulating temperature, humidity, watering regimes and the number of pollinators in commercial bumblebee hives. I tested for the occurrence of adaptive evolutionary trait shifts in a natural population of Matthiola tricuspidata with ancestors and descendants sampled 28 years apart.
The results show significant advancement of the flowering date in descendants compared to ancestors, consistent with adaptation of phenology to warmer and drier springs. Petal size and anther-stigma distance increased in the present-day environment, but did not show significant differences between ancestors and descendants, which is consistent with plastic trait shifts towards increased pollinator attractivity but not adaptive evolution. In line with this, pollinator-mediated pollination was more efficient in descendants compared to ancestors, and autonomous pollination was more efficient in ancestors compared to descendants. Altogether, these results suggest an evolution towards increased pollinator attractivity and outcrossing in the descendants. (Figure 1).
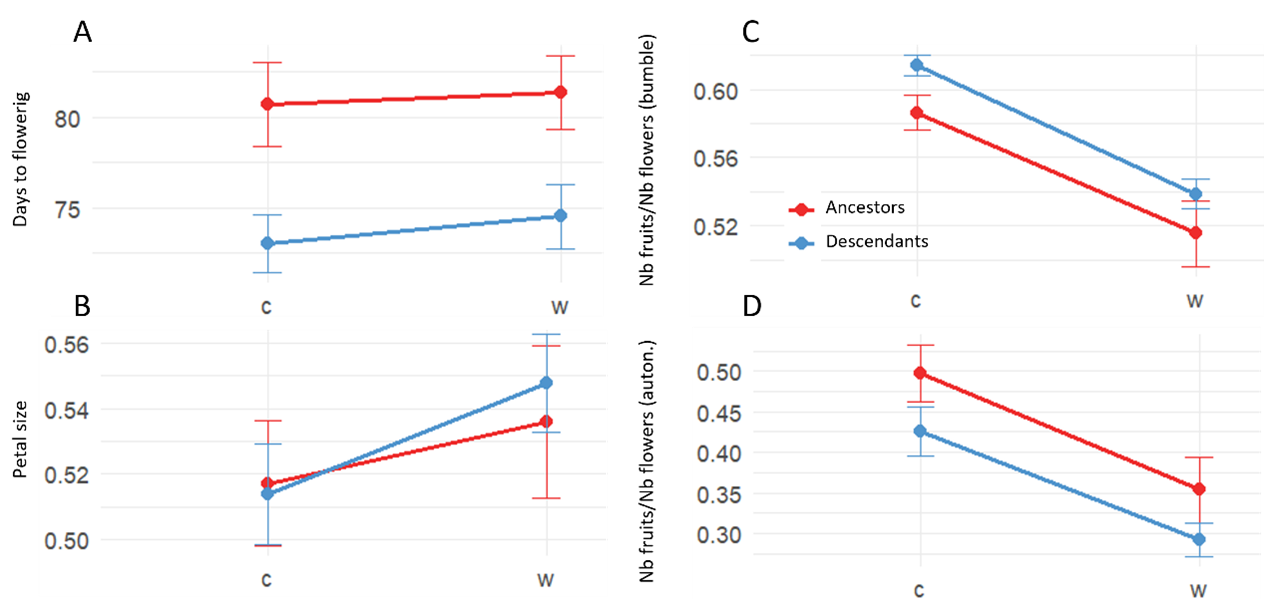
Resurrection ecology and genomics
Evolutionary trait shifts are not necessarily adaptive – they can be the results of stochastic processes such as drift, founder effects, migration, or even extinction-recolonization dynamics. To test whether phenotypic trait shifts revealed with the resurrection approach are truly the result of selective processes, I investigated neutral genetic variation assessed with ddRAD SNPs between ancestors and descendants in two natural populations (one sampled in Belgium and one in Poland) of Centaurium erythraea sampled 17 and 24 years apart.
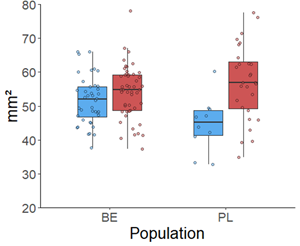
The descendants in both populations showed delayed flowering phenology, consistent with adaptation to avoid prolonged summer drought. In both populations, descendants had larger daily floral displays, larger petals (Figure 2), whereas the anther-stigma distance remained unchanged. Altogether, the observations show an evolution to increased flower attractiveness that could maintain plant-pollinator interactions.
Pairwise FST estimates showed low differentiation between ancestors and descendants in Belgium (0.077), and high differentiation in Poland (0.292). This suggests the occurrence of major evolutionary or demographic changes in the Polish population since the sampling of ancestral genotypes (e.g. migration, extinction and recolonization) or possible high sampling bias in the past. This result illustrates a shortcoming of resurrection studies, namely the difficulty to disentangle the effects of natural selection from stochastic variation in allelic frequencies due to random evolutionary processes or sampling bias (Etterson et al. 2016). Testing for different evolutionary scenarios using approximate Bayesian computations (ABC) could bring insight into the population evolutionary history (Collin et al. 2021). This nevertheless requires genotyping neighboring populations to be able to infer the probability that the sampled descendant genotypes are direct offspring of the ancestral genotypes present in 2003 relative to the probability that the contemporary genotypes were introduced from another population in its vicinity.
Resurrection epigenomics
With the recent accumulation of empirical observation about the existence of transgenerational phenotypic plasticity and hereditary epigenetic variation (Richards et al. 2017), a new issue rises in the common garden studies of adaptive evolution – observed phenotypic trait shifts could be due to evolutionary changes that modify the allelic frequences of a population over time, or they could be the result of heritable epigenetic modifications which do not affect the population genetic composition. Ecological demethylation assays are a cost-effective method to test for heritable epigenetic variation. By generally decreasing methylation levels throughout the genome, if the phenotypic differences between ancestors and descendants still persist, it is then likely that the observed trait shifts are mediated by genetic rather than epigenetic differences.
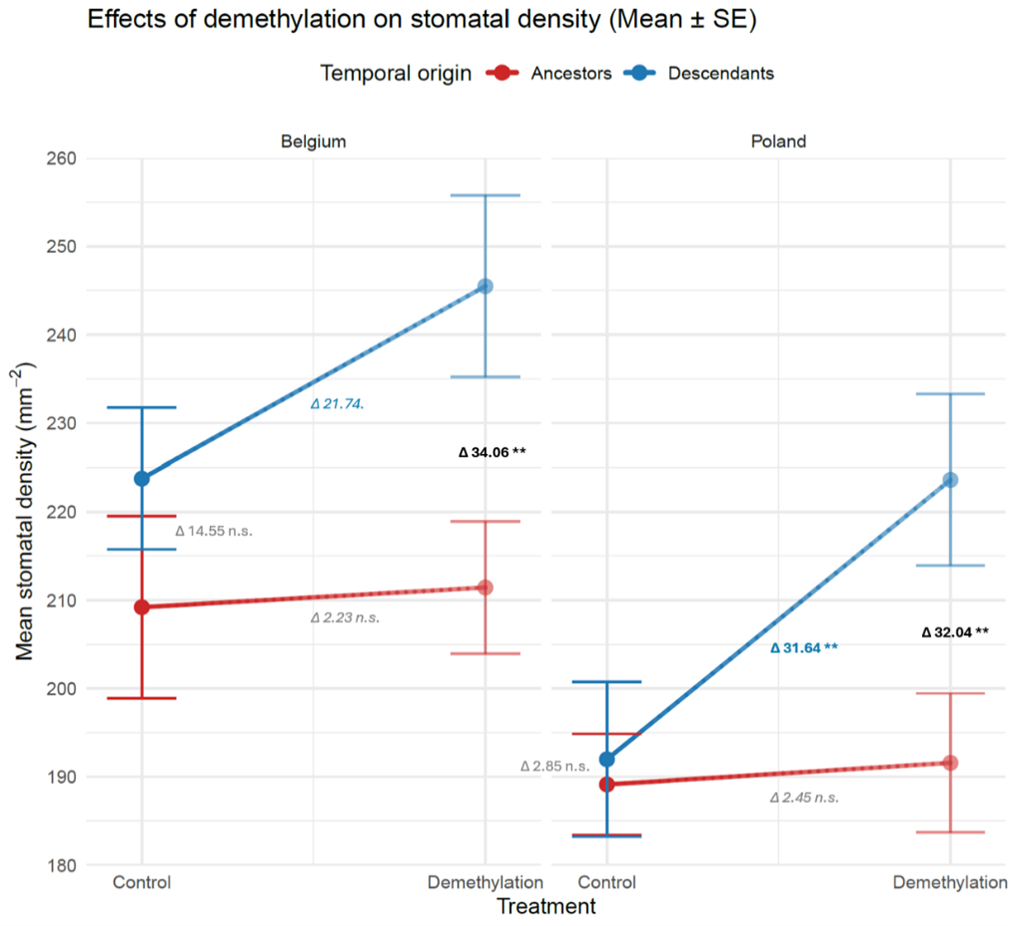
I implemented ecological demethylation assays in a common garden resurrection experiment to evaluate the effects of genetic and epigenetic variation on phenotypic trait shifts between ancestors and descendants un two populations of Centaurium erythraea. To my knowledge, this is the first resurrection epigenetics study in plants. The data confirms the phenotypic trait shifts between ancestors and descendants in a control environment without a demethylation agent. Applying a demethylation treatment results in strikingly different patterns depending on the measured traits. For some traits, demethylation reduces the differences between ancestors and descendants suggesting hereditary epigenetic variation (Figure 3). For others, demethylation has no effect, suggesting genetic regulation (Figure 4). Most surprisingly, in some traits, demethylation increases the differences between ancestors and descendants (Figure 5). This suggests that some genetic differences between ancestors and descendants were masked by epigenetic variation. Whether this pattern is due to epigenetic effects induced by the common environment, or hereditary epigenetic variation that masks genetic differences (canalization), remains to be investigated.
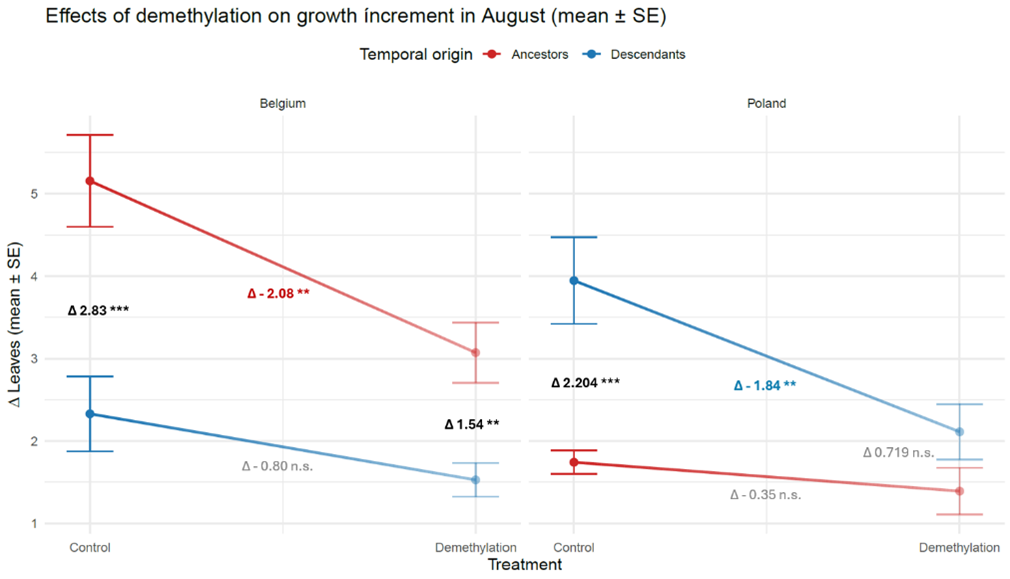
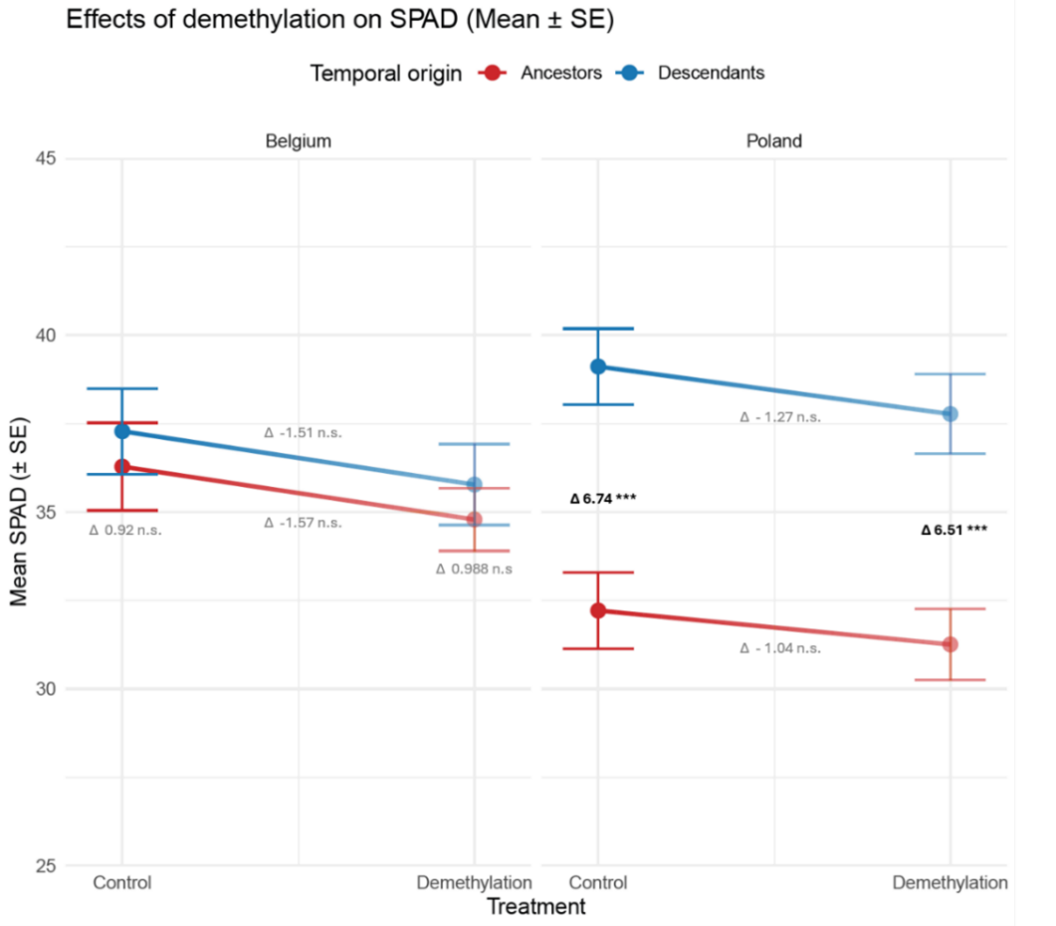
Conclusions
The resurrection method is an innovative way to study the occurrence of rapid adaptation in long-lived organisms by “instant” phenotypic comparisons of ancestral and descendant genotypes. Using this method, I show that arable weeds can rapidly adapt to fast-paced, human-mediated changes such as pollinator decline and climate change. A promising result is that the observed shifts in floral traits are consistent with evolution towards increased flower attractivity, which could help maintain plant-pollinator interactions despite global pollinator decline. While such adaptations are limited by plant physiology and resource availability, they still delay the decoupling of plant-pollinator interactions. This delay allows additional time for the implementation of conservations programs that will preserve plants and pollinators alike. These shifts are likely the outcome of adaptive evolutionary processes that can act on both the genetic and epigenetic composition of the population.
- Acoca-Pidolle, Samson, Perrine Gauthier, Louis Devresse, Antoine Deverge Merdrignac, Virginie Pons, and Pierre-Olivier Cheptou. 2024. “Ongoing Convergent Evolution of a Selfing Syndrome Threatens Plant–Pollinator Interactions.” New Phytologist 242 (2): 717–26. https://doi.org/10.1111/nph.19422.
- Brys, Rein, and Hans Jacquemyn. 2012. “Effects of Human-Mediated Pollinator Impoverishment on Floral Traits and Mating Patterns in a Short-Lived Herb: An Experimental Approach.” Functional Ecology 26 (1): 189–97. https://doi.org/10.1111/j.1365-2435.2011.01923.x.
- Cheptou, Pierre-olivier, Eric Imbert, and Michel Thomann. 2022. “Rapid Evolution of Selfing Syndrome Traits in Viola Arvensis Revealed by Resurrection Ecology.” American Journal of Botany 109 (11): 1838–46. https://doi.org/10.1002/ajb2.16028.
- Collin, François-David, Ghislain Durif, Louis Raynal, et al. 2021. “Extending Approximate Bayesian Computation with Supervised Machine Learning to Infer Demographic History from Genetic Polymorphisms Using DIYABC Random Forest.” Molecular Ecology Resources 21 (8): 2598–613. https://doi.org/10.1111/1755-0998.13413.
- Etterson, Julie R., Steven J. Franks, Susan J. Mazer, et al. 2016. “Project Baseline: An Unprecedented Resource to Study Plant Evolution across Space and Time.” In American Journal of Botany, vol. 103. no. 1. WILEY. https://doi.org/10.3732/ajb.1500313.
- Franks, S. J., E. Hamann, and A. E. Weis. 2018. “Using the Resurrection Approach to Understand Contemporary Evolution in Changing Environments. Evolutionary Applications 11: 17–28.”
- Richards, Christina L., Conchita Alonso, Claude Becker, et al. 2017. “Ecological Plant Epigenetics: Evidence from Model and Non-Model Species, and the Way Forward.” Ecology Letters 20 (12): 1576–90.
- Thomann, M., E. Imbert, and P. O. Cheptou. 2015. “Is Rapid Evolution of Reproductive Traits in Adonis Annua Consistent with Pollinator Decline?” In Acta Oecologica, vol. 69. ELSEVIER. https://doi.org/10.1016/j.actao.2015.10.007.
- Valencia-Montoya, Wendy A., Elodie Flaven, Juliette Pouzadoux, Eric Imbert, and Pierre-Olivier Cheptou. 2021. “Rapid Divergent Evolution of an Annual Plant across a Latitudinal Gradient Revealed by Seed Resurrection.” In Evolution, vol. 75. no. 11. WILEY. https://doi.org/10.1111/evo.14364.